Bioinformatics researcher uses Anvil AI to advance field and present at NeurIPS 2025
A research group from the University of Missouri recently used Purdue’s Anvil supercomputer to train a deep learning model to represent three-dimensional protein structures that are effective in downstream bioinformatics tasks. The new model not only achieved more accurate protein reconstructions than prior methods but also paves the way for future applications, thanks to the team's insistence on a fully open-source implementation and data access, which are currently lacking in the field. This research was presented at NeurIPS 2025—a renowned international machine learning and computational neuroscience conference—as part of the AI4Science Workshop.
Dong Xu was a Curators’ Distinguished Professor in the Department of Electrical Engineering and Computer Science at the University of Missouri. He joined the Health Informatics Institute at the University of South Florida, as a Professor this year. He is the head of the Digital Biology Lab, a research group focused on the intersection of bioinformatics and artificial intelligence (AI). Recently, Xu and his team decided to tackle a major challenge in the field of computational biology—understanding and predicting protein structures.
Proteins are the building blocks of life, serving many crucial roles in biochemistry. Each protein has its own unique three-dimensional (3D) structure that is intricately tied to its function. Scientists estimate that there are more than 2 million different protein structures. However, not all of these are yet known, much less understood. Being able to accurately predict different protein shapes is the key to unlocking major discoveries in pharmaceutical design, disease comprehension, and biotechnology.
Current research 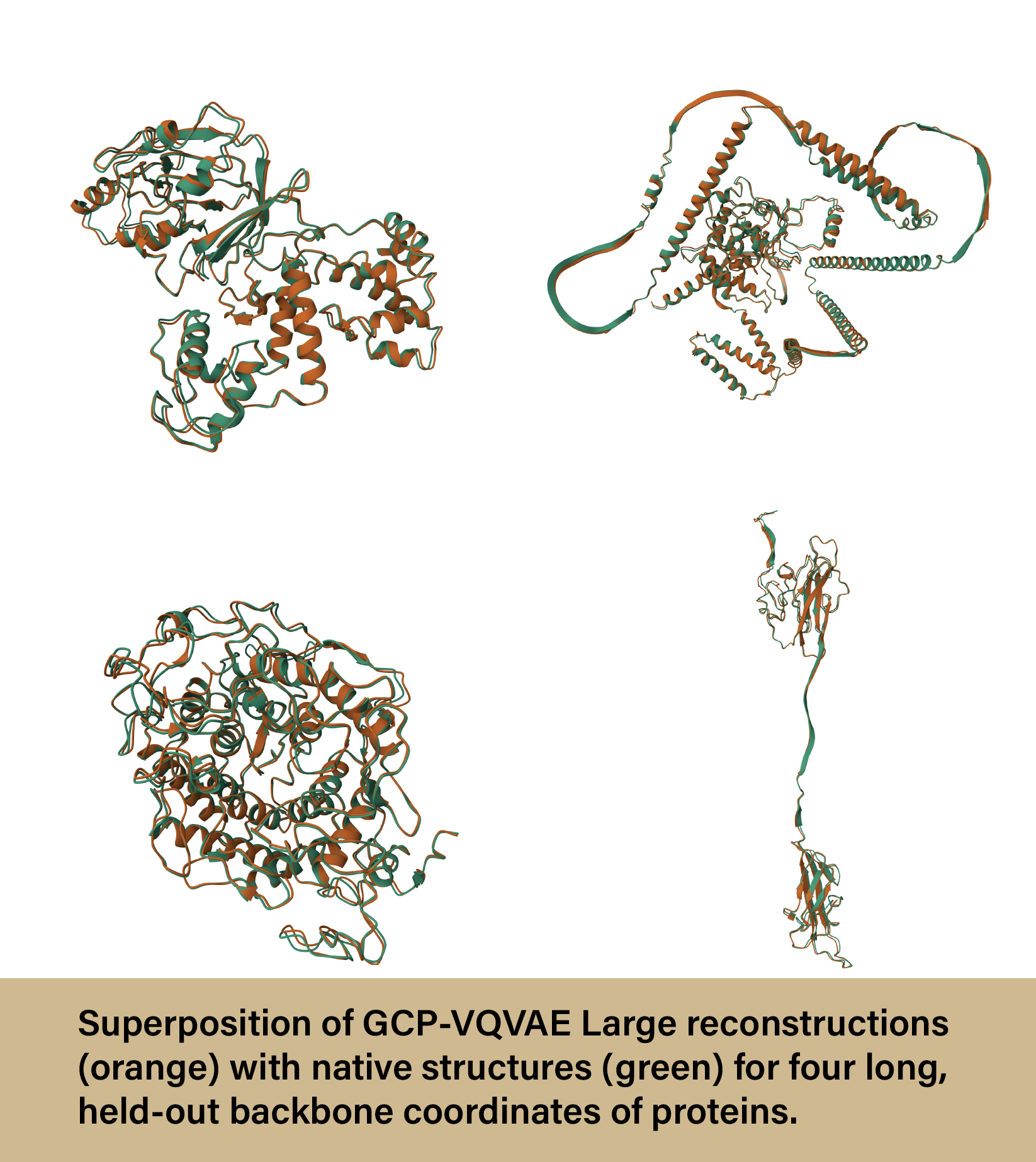 suggests that protein structure follows a linear logic. Much like natural language, which conforms to rules of grammar, proteins seem to have their own rules that instruct how their 3D structures are formed. Knowing this, scientists can try to leverage the underlying “grammar” along with AI methods to develop a language of 3D geometry. This language can then be used to reconstruct and even predict 3D protein structures. Utilizing high-performance computing (HPC) resources to simulate and predict protein structures is not new to the field of bioinformatics, but despite recent advances, creating representations of the 3D geometry of proteins that can be used in generative modeling and downstream AI tasks has proven elusive. Xu and his lab decided to use the Anvil supercomputer to address this challenge.
suggests that protein structure follows a linear logic. Much like natural language, which conforms to rules of grammar, proteins seem to have their own rules that instruct how their 3D structures are formed. Knowing this, scientists can try to leverage the underlying “grammar” along with AI methods to develop a language of 3D geometry. This language can then be used to reconstruct and even predict 3D protein structures. Utilizing high-performance computing (HPC) resources to simulate and predict protein structures is not new to the field of bioinformatics, but despite recent advances, creating representations of the 3D geometry of proteins that can be used in generative modeling and downstream AI tasks has proven elusive. Xu and his lab decided to use the Anvil supercomputer to address this challenge.
“For this project, we used Anvil to train a deep learning model to represent protein structures,” says Xu. “So if you think about a protein, it consists of amino acids—they form these chains that create a three-dimensional structure. You have probably heard of AlphaFold, which can predict the structure very well. But in order to really study the massive structures, either experimentally generated or computationally predicted, it requires a good representation of protein structure, one that can be used in an effective way to do downstream tasks. So we needed to develop these representations.”
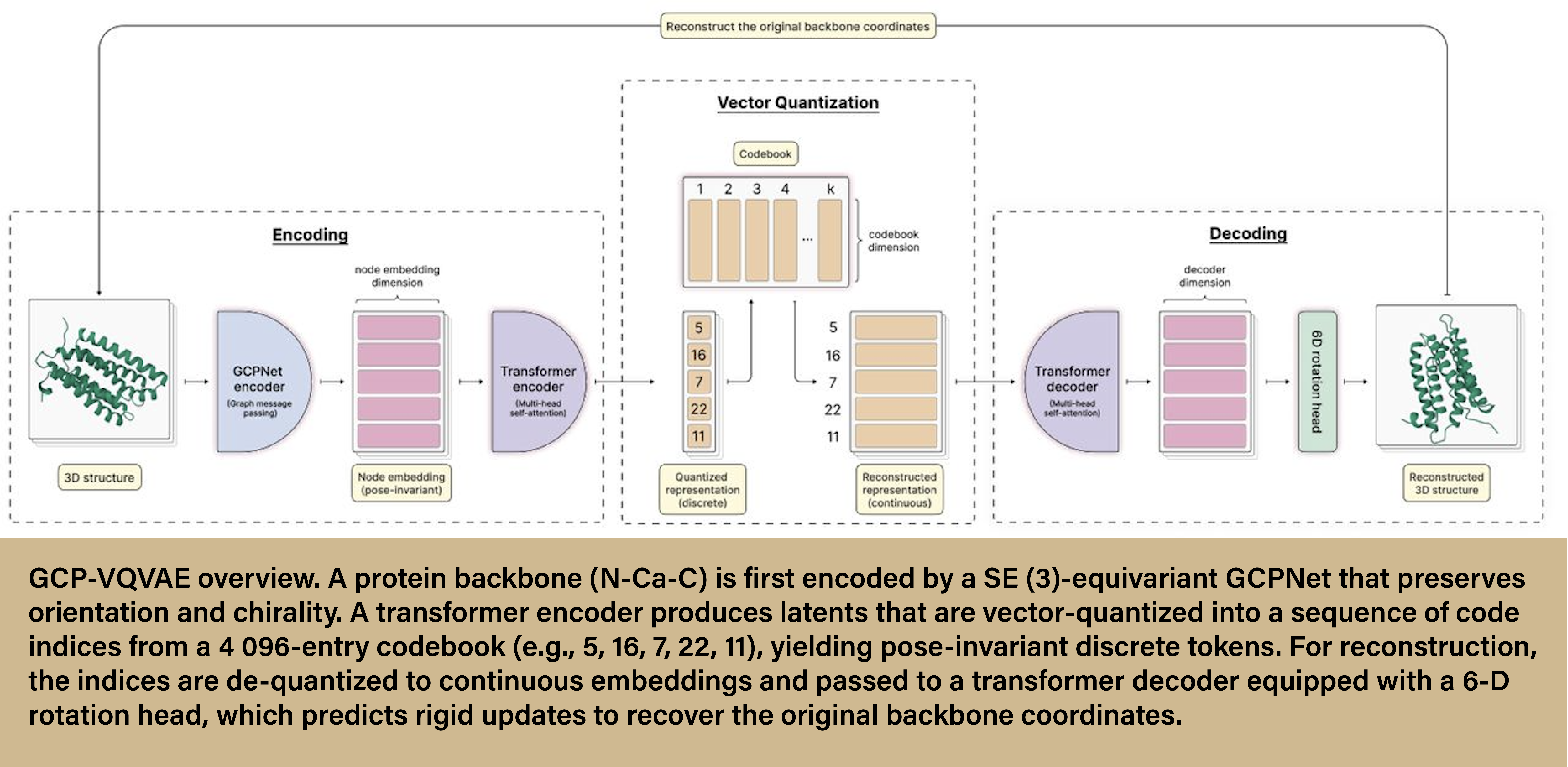
Through their NAIRR allocation, Xu and his team, especially his Ph.D. student Mahdi Pourmirzaei, had access to Anvil’s H100 GPUs, which they used to develop, train, and test GCP-VQVAE (geometry complete-vector-quantized variational autoencoder) on a corpus of 24 million monomer protein backbone structures. The team’s results were overwhelmingly positive.
GCP-VQVAE produced more accurate protein reconstructions than other deep learning models, maintaining proper orientation and chirality—a difficult task to achieve while still retaining accuracy. The model also demonstrated robust performance to unseen proteins, meaning it could produce protein structures that were not included as part of the training data. Lastly, the GCP-VQVAE model proved to be much more efficient than other models, achieving approximately 408x and 530x lower end-to-end latency. The best part? GCP-VQVAE is fully open-sourced with transparent training and evaluations. Any researcher can utilize the model, see how it was trained, and compare it with other VQVAEs.
To accomplish this, Xu’s lab relied on the advanced GPUs provided by Anvil. The group initially had access to A100 GPUs via their original NAIRR allocation. After starting the project, Xu realized they needed to work at a faster pace and reached out to the Anvil support team. The Anvil team convinced Xu to trade his A100 allocation hours for H100 hours, using Anvil’s Service Unit Swap feature. Although the swap comes at a 2:1 ratio (two A100 hours for one H100 hour), Xu noted that the results were well worth the trade. Not only are the H100s more powerful and efficient than the A100s, but the H100 queue on Anvil is significantly shorter, allowing Xu to run his jobs in a more condensed time frame. All of this led to a much faster time-to-science for the group.
“We initially had an allocation of A100s,” says Xu, “but there were a lot of jobs, so people have to wait. The Anvil staff suggested we trade to the H100s, and when we did that, we didn’t have a queue; the jobs ran immediately. So we got more computing power from the trade, and we didn’t have to wait. It worked really, really well.”
Xu continued, “Anvil also provided top-notch customer service. They gave us a lot of help, which I really appreciate. We had meetings with them, they answered emails very quickly, and they gave a lot of guidance for our students and postdocs.
Overall, Xu and his research lab were immensely pleased with the Anvil supercomputer. He noted that thanks to NAIRR and Anvil, his team was able to produce multiple papers and present at major conferences, such as the AI4Science Workshop at NeurIPS 2025. Aside from the NeurIPS presentation, Anvil helped the team to produce a preprint in bioRxiv and a publication in Computational and Structural Biotechnology Journal.
“Resources like Anvil, they are really valuable to us,” says Xu. “We ran a lot of jobs, and have already published papers acknowledging the allocation; it’s extremely important to us. Without these resources, we wouldn’t be able to do much.”
For more information on Xu’s research, please visit: Dong Xu’s Google Scholar Page.
To learn more about High-Performance Computing and how it can help you, please visit our “Why HPC?” page.
Anvil is one of Purdue University’s most powerful supercomputers, providing researchers from diverse backgrounds with advanced computing capabilities. Built through a $10 million system acquisition grant from the National Science Foundation (NSF), Anvil supports scientific discovery by providing resources through the NSF’s Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS), a program that serves tens of thousands of researchers across the United States. Anvil also supports advanced artificial intelligence research as an official resource provider of the National Artificial Intelligence Research Resource (NAIRR) Pilot.
Researchers may request access to Anvil via the ACCESS allocations process or through the NAIRR allocations process. More information about Anvil is available on Purdue’s Anvil website. Anyone with questions should contact anvil@purdue.edu. Anvil is funded under NSF award No. 2005632.
Written by: Jonathan Poole, poole43@purdue.edu
